Research InterestChanges in physiological conditions, for example, oxygen and nutrient deprivation, viral infection, oncogenic mutations, exposure to UV irradiation, and suboptimal growth environment, pose threats to cells. Cells have developed sophisticated adaptive mechanisms to respond to external and internal stresses. Cell signalling events, such as phosphorylation and changes of cellular calcium levels, provide immediate responses. Post-transcriptional regulation, mainly mRNA biogenesis and control of translation, functions at an intermediate time-scale. In contrast, regulation of gene expression is essential for the slower, long-term adaptation and recovery phases. Stress response is a conserved mechanism for all organisms and the dysregulation of which leads to many diseases.
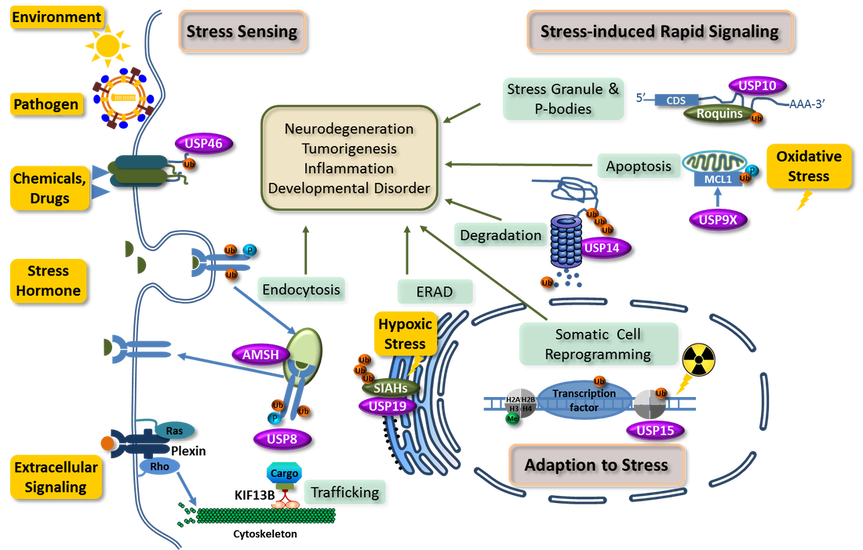
Ubiquitination, the second most prevalent post-translational modification in eukaryotic cells, plays a vital role in all aspects of stress response. The diagram below summarizes the protein systems we have worked on in the last decade and their connection to the stress response processes. We continue to explore the structure, function, and regulation of protein systems important for stress response . |
The Three Pillars of Our Research Program
TopicsUbiquitination
Ubiquitin (Ub) is a small, highly conserved, regulatory protein consists of 76 amino acids. Ubiquitination is the reversible covalent conjugation of Ub to a substrate protein through an isopeptide between the C-terminal carboxyl group of the Ub with the amine group of a lysine side chain (exceptions exist). Ubiquitination is catalyzed by a three-enzyme cascade that comprises of E1 activating enzyme, E2 conjugating enzyme and E3 ligase, and the Ub tag can be removed by a family of deubiquitinases (DUBs). Ub has seven lysines, plus the first methione, that can be further ubiquitinated to form a poly-Ub chain of different linkages. Poly-Ub chain linkage dictates the fate of the substrate protein. |
TechnologiesStructural biology
Structural biology studies the molecular structure and dynamics of biomacromolecules using biophysical techniques. Scientists mainly use three technologies to determine the atomic details of biomolecules: nuclear magnetic resonance (NMR) spectroscopy, X-ray crystallography, and the latest single-particle cryogenic electron microscopy. But other biophysical techniques, such as small-angle X-ray scattering (SAXS), hydrogen/deuterium exchange mass spectroscopy (H/D-MS), dynamic light scattering, can also be used to answer specific questions of molecular structure and dynamics. It is crucial that we choose the right tools to answer the biological question at hand. |
ApplicationsBasic research
Macromolecular interactions, aka "molecular recognition," are often specific and highly dynamic, which is essential for the tight regulation of biological processes. Elucidating the molecular details that mediate protein-compound, protein-protein and protein-nucleic acid interactions are crucial to understanding the fundamental mechanisms of cell signalling, DNA damage repair, host-pathogen interaction, drug targeting, and many others. Drug discovery Modern target-based drug discovery is a complicated process involving many stages. Target validation is a critical step with the highest attrition rate. High-quality tool molecules, such as chemical probes, or ubiquitin variants, are the "holy grail" for mimicking pharmacological interference to validate the therapeutic value of the cognate targets. Structural biology plays a vital role in developing these tools. |
Copyright © 2019 Yufeng Tong. All rights reserved.